Estimates the predictive performance of predictions under interventions, by forming a weighted pseudopopulation in which every subject was assigned the treatment of interest.
Usage
ip_score(
object,
data,
outcome,
treatment_formula,
treatment_of_interest,
metrics = c("auc", "brier", "oeratio", "calplot"),
time_horizon,
cens_model = "KM",
cens_formula = ~1,
null_model = TRUE,
stable_iptw = FALSE,
bootstrap = 0,
bootstrap_progress = TRUE,
iptw,
ipcw,
quiet = FALSE,
strip_ipt_models = TRUE
)Arguments
- object
One of the following three options to be validated:
a numeric vector, corresponding to risk predictions under intervention of interest.
a glm or coxph model, capable of estimating risks under intervention of interest. See details.
a (named) list, with one or more of the previous 2 options, for validating and comparing multiple models at once.
- data
A data.frame containing the observed outcome, assigned treatment, and necessary confounders for the validation of object.
- outcome
The outcome, to be evaluated within data. This could either be the name of a numeric/logical column in data, or a Surv object for time-to-event data, e.g. Surv(time, status), if time and status are columns in data.
- treatment_formula
A formula which identifies the treatment (left hand side) and the confounders (right hand side) in the data. E.g. A ~ L. The treatment can be either binary (0/1) or a categorical factor. The confounders are used to estimate the inverse probability of treatment weights (IPTW) model. The IPTW can also be specified themselves using the iptw argument, in which case the right hand side of this formula is ignored (the left hand side must still identify the treatment, i.e. A ~ 1).
- treatment_of_interest
A treatment level for which the interventional performance measures should be evaluated.
- metrics
A character vector specifying which performance metrics to be computed. Options are c("auc", "brier", "oeratio", "calplot"). See details.
- time_horizon
For time to event data, the prediction horizon of interest.
- cens_model
Model for estimating inverse probability of censored weights (IPCW). Methods currently implemented are Kaplan-Meier ("KM") or Cox ("cox"), both applied to the censored times. KM is only supported when the right hand side of cens_formula is 1.
- cens_formula
Formula for which the r.h.s. determines the censoring probabilities. I.e. ~ x1 + x2.
- null_model
If TRUE fit a risk prediction model which ignores the covariates and predicts the same value for all subjects. The model is fitted using the data in which all subjects are 'counterfactually' assigned the treatment of interest (using the IPTW, as estimated using the treatment_formula or as given by the iptw argument). For time-to-event outcomes, the subjects are also 'counterfactually' uncensored (using the IPCW, as estimated using the cens_formula, or as given by the ipcw argument).
- stable_iptw
if TRUE, estimate stabilized IPT-weights. Does not influence the metrics. See details.
- bootstrap
If this is an integer greater than 0, this indicates the number of bootstrap iterations, to compute 95% confidence intervals around the performance metrics.
- bootstrap_progress
if set to TRUE, print a progress bar indicating the progress of bootstrap procedure.
- iptw
A numeric vector, containing the inverse probability of treatment weights. These are normally computed using the treatment_formula, but they can be specified directly via this argument. A function can also be specified, which takes as input 'data' and returns a numeric vector of IPTW weights. See details.
- ipcw
A numeric vector, containing the inverse probability of censoring weights. These are normally computed using the cens_formula, but they can be specified directly via this argument. A function can also be specified, which takes as input 'data' and returns a numeric vector of IPCW weights. See details.
- quiet
If set to TRUE, don't print assumptions.
- strip_ipt_models
If set to TRUE (default), the models for the IPT and IPC-weights are stripped of unnecessary data. Set to FALSE if you plan to do extensive diagnostics on the fitted IPT/IPC models. The resulting `ip_score` object will use quite a lot more memory.
Value
An object of class `ip_score`, for which the `print()` and `plot()` methods are implemented. The object is a nested list containing:
`$score`, which contains the predictive performance in the pseudopopulation dataset.
`$bootstrap`, if applicable, the 95% confidence intervals of the performance metrics, and the performance metrics for each individual bootstrap iteration.
`$outcome`, the observed outcome of the original dataset.
`$treatment`, the observed outcome of the original dataset.
`$predictions`, the predictions to be evaluated, i.e. the probability of event for each patient, had their treatment been set to treatment_of_interest.
`$ipt`, method, model and inverse probability of treatment weights (IPTW). These are NA for patients that are not in the pseudopopulation.
`$ipc`, method, model and inverse probability of censoring weights (IPCW). These are NA for patients that were censored.
`$pseudopop`, binary vector indicating which subjects of the original population were in the pseudopopulation, by following the treatment of interest and remaining uncensored, if applicable.
The print method summarizes the results and if (quiet = FALSE), prints the assumptions required for valid inference.
Details
When supplying a glm or coxph model, the model should be able to estimate risks under the intervention of interest. This could be done in two ways: the model does not have a treatment covariate, and always estimates the risk under intervention of interest. Alternatively, the model has a covariate for treatment. This function then automatically estimates the risk under the treatment of interest for all subjects, even if they were assigned alternative treatment.
All performance metrics are computed on the weighted population in which
every subject was counterfactually assigned treatment of interest. auc
is area under the (ROC) curve. Brier score is defined as 1 / sum(iptw)
sum(predictions_i - outcome_i)^2. Scaled brier score is also available
(metrics = "scaled_brier"). For the O/E ratio, the numerator
(observed) is the weighted fraction of events in the pseudopopulation, and
the denominator (expected) is the unweighted mean of risk estimates of the
original unweighted population. The calplot option generates a
calibration plot, with default 8 subgroups. More/less subgroups can be
specified by appending calplot with a number indicating the number of
subgroups, e.g. metrics = calplot10 for 10 subgroups.
For the null model, the O/E ratio and the scaled Brier score, the mean predicted risk under the treatment of interest is required. This is computed by the weighted mean in the ('counterfactually' uncensored) pseudopopulation. For survival data, this null prediction could theoretically also be computed with a weighted Kaplan-Meier, which is supposed to be more efficient, but computationally a lot slower. Both methods are valid.
Stabilized IPT-weigths are computed by estimating a null model for treatment.
E.g. weights are P(A = a) / P(A = a | L = l), if the given
treatment_formula is A ~ L. In a pseudopopulation in which
everybody's treatment was set to a certain level `a`, the numerator of this
expression is constant. The resulting performance metrics are not influenced
by multiplication of weights with a constant.
Bootstrap is not possible when manually specifiying the IPTW/IPCW as numeric vectors. If specifying a function that computes the ITPW/IPCW given data, it is possible. The given function will be called on each bootstrapped dataset to compute the 95% CI of the performance metrics. More advanced techniques, such as thresholding extreme IP weights, can be implemented this way. The censoring weight returned should be the 1 / probability of remaining uncensored at the time horizon, or at their event time, whichever happens first.
References
Keogh RH, Van Geloven N. Prediction Under Interventions: Evaluation of Counterfactual Performance Using Longitudinal Observational Data. Epidemiology. 2024;35(3):329-339.
Boyer CB, Dahabreh IJ, Steingrimsson JA. Estimating and Evaluating Counterfactual Prediction Models. Statistics in Medicine. 2025;44(23-24):e70287.
Pajouheshnia R, Peelen LM, Moons KGM, Reitsma JB, Groenwold RHH. Accounting for treatment use when validating a prognostic model: a simulation study. BMC Medical Research Methodology. 2017;17(1):103.
Examples
n <- 1000
data <- data.frame(L = rnorm(n), P = rnorm(n))
data$A <- rbinom(n, 1, plogis(data$L))
data$Y <- rbinom(n, 1, plogis(0.1 + 0.5*data$L + 0.7*data$P - 2*data$A))
random <- runif(n, 0, 1)
model <- glm(Y ~ A + P, data = data, family = "binomial")
naive_perfect <- data$Y
score <- ip_score(
object = list("ran" = random, "mod" = model, "per" = naive_perfect),
data = data,
outcome = Y,
treatment_formula = A ~ L,
treatment_of_interest = 0,
)
print(score)
#> Estimation of the performance of the prediction model in a
#> pseudopopulation where everyone's treatment A was set to 0.
#> The pseudopopulation ($pseudopop) is constructed from 494 (49.4%)
#> subjects who indeed received treatment level 0. These subjects are
#> reweighted to represent the full target population under a hypothetical
#> intervention in which everyone received this treatment level.
#> The following assumptions must be satisfied for correct inference:
#>
#> Causal assumptions:
#>
#> - Conditional exchangeability: after adjustment for the covariates used
#> to construct the inverse probability of treatment weights (IPTW), i.e.,
#> {L}, there are no unmeasured confounders of treatment assignment and
#> outcome.
#> - Conditional positivity: the probability of receiving treatment level
#> 0 should be greated than zero for each combination of the variables {L}
#> that is observed in the full population. The distribution of
#> IPT-weights can be assessed with $ipt$weights[$pseudopop$ids].
#> - Consistency: the observed outcome under the received treatment equals
#> the potential outcome under that treatment. This includes the
#> assumption of no interference between subjects.
#>
#> Modeling assumptions:
#>
#> - Correctly specified propensity model. Estimated treatment model is
#> logit(A) = 0.02 + 0.98*L. See also $ipt$model.
#>
#> Performance estimates:
#>
#> model auc brier oeratio
#> null model 0.500 0.249 1.00
#> ran 0.544 0.310 1.04
#> mod 0.668 0.232 1.17
#> per 1.000 0.000 1.61
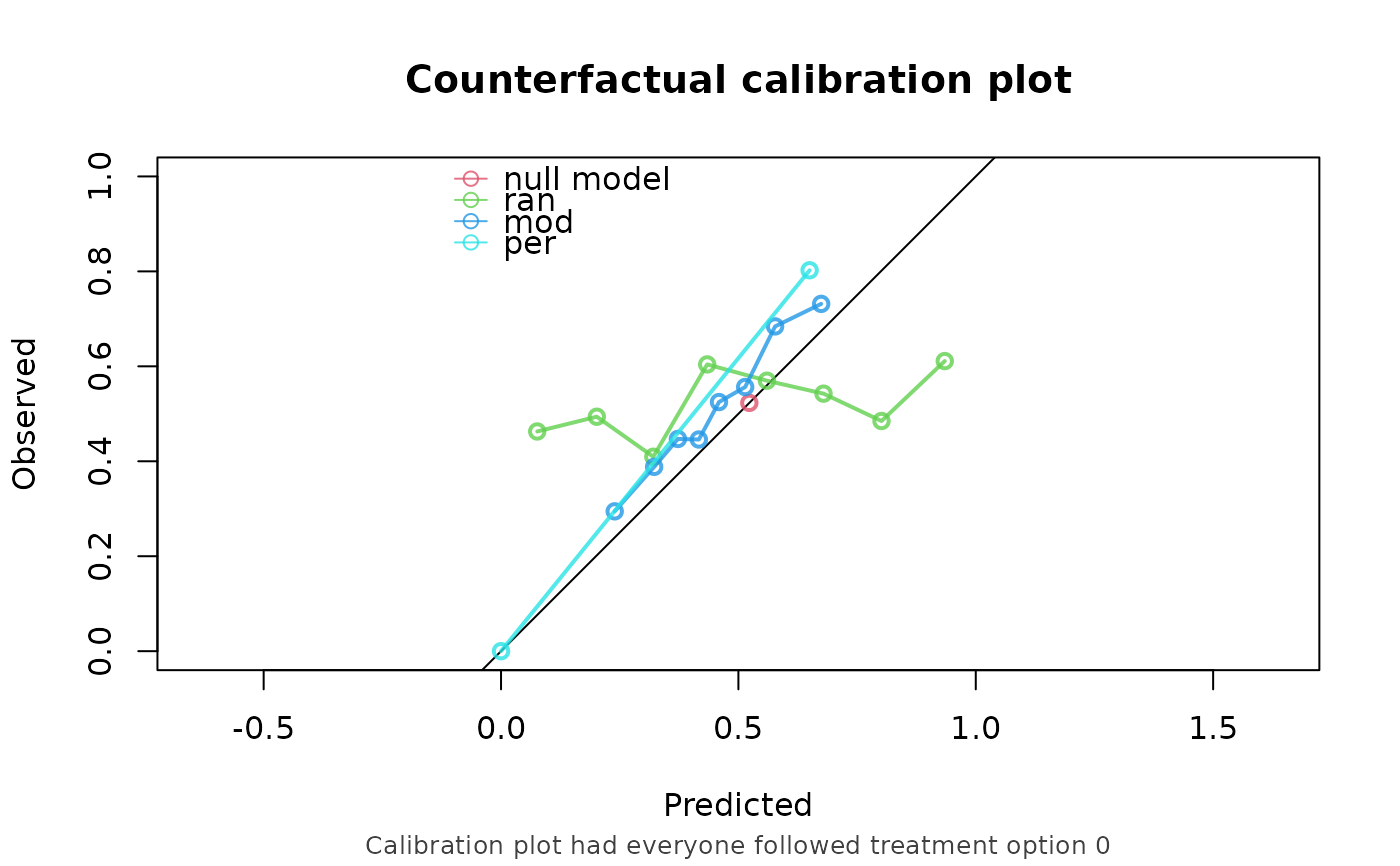 plot(score)
plot(score)